GGE: A Standardized Framework for Evaluating Gene Expression Generative Models
Paper: Accepted at the Gen2 Workshop at ICLR 2026
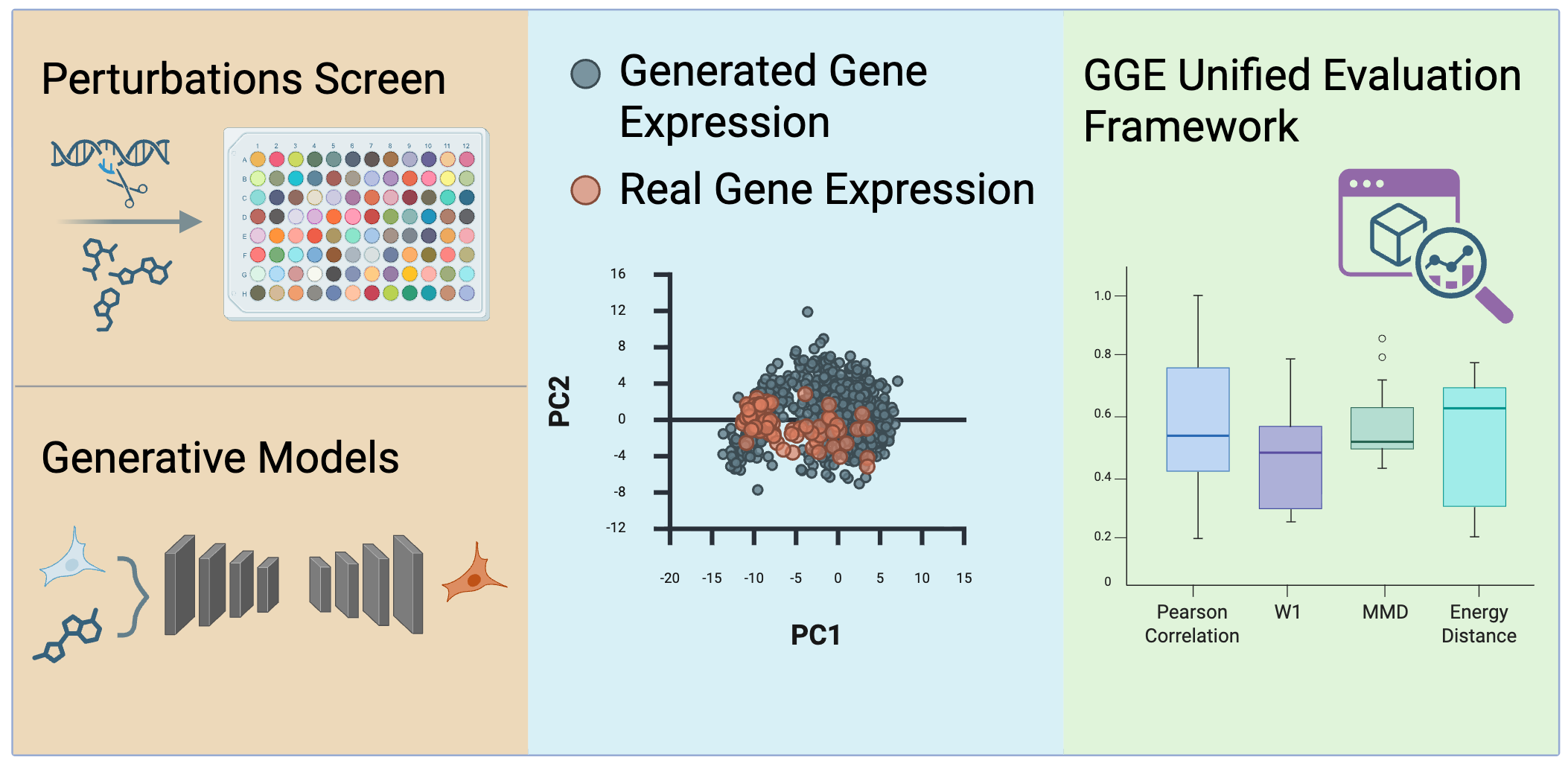
Comprehensive, standardized evaluation of generated gene expression data.
GGE (Generated Genetic Expression Evaluator) addresses the urgent need for standardized evaluation in single-cell gene expression generative models. Current practices suffer from inconsistent metric implementations, incomparable hyperparameter choices, and lack of biologically-grounded metrics. GGE provides:
- Comprehensive suite of distributional metrics with explicit computation space options
- Biologically-motivated evaluation through DEG-focused analysis with perturbation-effect correlation
- Standardized reporting for reproducible benchmarking
Full Documentation with API tutorials can be found here
Key Features
- Per-metric space configuration (raw, PCA, DEG)
- Perturbation-effect correlation (Paper Eq. 1)
- Configurable DEG thresholds
- GPU (CUDA) and Apple MPS acceleration
- Per-gene and aggregate metrics
- Publication-quality visualizations (static and interactive)
- Simple Python API and CLI
- Mixed-space evaluation with
evaluate_lazy()
Metrics
All metrics are computed per-gene (returning a vector) and aggregated:
| Metric | Description | Direction |
|---|---|---|
| Pearson Correlation | Linear correlation between expression profiles | Higher is better |
| Spearman Correlation | Rank correlation (robust to outliers) | Higher is better |
| R² | Coefficient of determination | Higher is better |
| Perturbation-Effect Correlation | Correlation on (real - ctrl) vs (gen - ctrl) | Higher is better |
| MSE | Mean Squared Error | Lower is better |
| Wasserstein-1 | Earth Mover's Distance (L1) | Lower is better |
| Wasserstein-2 | Sinkhorn-regularized OT | Lower is better |
| MMD | Maximum Mean Discrepancy (RBF kernel) | Lower is better |
| Energy Distance | Statistical potential energy | Lower is better |
Visualizations
- Boxplots and violin plots for metric distributions
- Radar plots for multi-metric comparison
- Scatter plots for real vs generated expression
- Embedding plots (PCA/UMAP) for real vs generated data
- Heatmaps for per-gene metric values
- Interactive Plotly plots with density overlays and metadata coloring
Computation Spaces
GGE treats computation space as a first-class parameter (see Paper Section 3.3):
| Space | Description | When to Use |
|---|---|---|
| Raw Gene Space | Full ~5,000–20,000 gene dimensions | Gene-level interpretability needed |
| PCA Space | Reduced k-dimensional space (default: 50) | Primary distributional metrics |
| DEG Space | Restricted to differentially expressed genes | Biologically-targeted evaluation |
Recommendation: Use multi-space evaluation—PCA-50 for distributional metrics, DEG for biological focus.
Installation
The package includes GPU-accelerated metrics via geomloss, which automatically falls back to CPU if no GPU is available.
Quick Start
Python API
from gge import evaluate
# From file paths
results = evaluate(
real_data="real_data.h5ad",
generated_data="generated_data.h5ad",
condition_columns=["perturbation", "cell_type"],
split_column="split", # Optional: for train/test
output_dir="evaluation_output/"
)
# From AnnData objects
import scanpy as sc
real_adata = sc.read_h5ad("real_data.h5ad")
generated_adata = sc.read_h5ad("generated_data.h5ad")
results = evaluate(
real_data=real_adata,
generated_data=generated_adata,
condition_columns=["perturbation"],
)
# Access results
print(results.summary())
# Get metric for specific split
test_results = results.get_split("test")
for condition, cond_result in test_results.conditions.items():
print(f"{condition}: Pearson={cond_result.get_metric_value('pearson'):.3f}")
Contributing
Contributions are welcome! Please feel free to submit a pull request or open an issue.
Citation
If you use GGE in your research, please cite our paper:
@misc{rubbi2026gge,
title = {A Standardized Framework for Evaluating Gene Expression Generative Models},
author = {Rubbi, Andrea and Di Francesco, Andrea Giuseppe and Lotfollahi, Mohammad and Liò, Pietro},
year = {2026},
note = {Presented at the GenAI in Genomics Workshop at ICLR 2026},
url = {https://genai-in-genomics.github.io/}
}
or the software
@software{rubbi2026gge,
author = {Rubbi, Andrea},
title = {GGE: Generated Genetic Expression Evaluator},
year = {2026},
url = {https://github.com/AndreaRubbi/GGE}
}
License
This project is licensed under the MIT License. See the LICENSE file for details.


